10.3.2—
Translation of mRNA
10.3.2.1—
Initiation of the Polypeptide Chain
Initiation of protein synthesis on plant cytoplasmic ribosomes starts with the assembly of a mRNA•40s subunit complex which requires ATP for its formation and this step is followed by the addition of methionyl-tRNAi to form a mRNA•40s subunit•methionyl-tRNAi complex. The process requires GTP and at least two initiation factors (Weeks et al., 1972; Marcus et al., 1973). Initiation is completed by the addition of the 60s subunit and the 80s ribosome so formed is then able to accept the next aminoacyl-tRNA, thus starting the elongation process. The tRNAi Met is a special tRNA charged with unformylated methionine, which can enter the so-called 'P' site of the ribosome complex and base-pair with the initiator codon, AUG or GUG in the mRNA. Although mRNAs are translated from the 5'® 3' end, evidence from sequence studies of bacteriophage messages shows that the initiator triplet is not at the 5' terminus, but is set a considerable number of nucleotides in. For example, the first initiator codon of bacteriophage R17 RNA is preceded by 91 nucleotides which are not translated (Adams & Cory, 1970). In the case of polycistronic messengers, one or more initiator triplets will occur intramolecularly.
Evidence that plants have a special type of initiator tRNA, tRNAi Met , comes from the wheatgerm system (Leis & Keller, 1970; Marcus et al., 1970a; Tarrago et al., 1970; Ghosh et al., 1971), and from Vicia faba (Yarwood et al., 1971a),
where it has been shown that two major and one minor tRNAMet species are present. The minor and one of the major tRNAMet species function as chain initiators as shown by AUG-dependent binding (Tarrago et al., 1970; Yarwood et al., 1971a), and by N-terminal analyses of either the product of tobacco mosaic viral RNA-directed (Marcus et al., 1970a) or poly-AUG, poly-GU and endogenous messenger-directed incorporation (Yarwood et al., 1971a). Neither of the major tRNAMet species from either wheat or beans is formylated, in contrast to the initiator tRNAiMet of prokaryotes, and it is presumed that these are the cytoplasmic tRNAMet s for initiator and internal methionine residues. The minor, formylatable tRNAMet , is presumed to be the initiator of chloroplast protein synthesis (see chapter 11)
The hydrolysis of ATP precedes the formation of the first peptide bond and may be required for recycling of the methionyl-tRNAi binding factor. The requirement for ATP has also been demonstrated in the rabbit globin synthesizing system (Schreier & Staehelin, 1973), but not in those of prokaryotes. The order in which the initiator tRNA and mRNA bind to the small subunit in mammalian and prokaryote systems is in dispute; the order suggested above for wheatgerm by Marcus and his coworkers, therefore, whilst agreeing with some workers on those systems disagrees in particular with the suggestion of Legon et al., (1973) and Schreier and Staehelin (1973) that the initiator binds prior to the mRNA in rabbit globin synthesis.
Knowledge of the two plant initiator factors involved in binding methionyl-tRNAi and mRNA is much less complete than for the similar factors in bacterial and mammalian systems; details of the latter are, therefore, relevant. In bacteria there are three factors, IF-1, IF-2 and IF-3 involved in the initiation process. Factor IF-1 has a molecular weight of 9,400 daltons and is the most basic of the three proteins. Factor IF-2 has been shown to consist of two sub-components, both active, with molecular weights of 9 x 103 –100 × 103 daltons and 80 × 103 daltons although the latter is probably derived from the former by proteolysis during purification. Factor IF-3 is a protein or group of proteins with molecular weight(s) of 21.5 × 103 –23.5 × 103 daltons (Grunberg-Manago et al., 1973). The three factors attach to the small subunit; IF-2 binds GTP and formyl-methionyl-RNA, while IF-3 binds mRNA. Factor IF-I increases the affinity of factors IF-2 and IF-3 for the 30s subunit, and is also necessary for the release of IF-2 from the ribosome.
The order of the attachment of the different factors to the small subunit and the order of the binding of mRNA and acylated initiator tRNA, is still a matter of debate. Some results suggest that IF-1 is attached before IF-2 and IF-3, whilst others indicate that it can only be attached after. Similarly, it is possible that an unstable intermediary complex consisting of formyl-methionyl-tRNAi •30s subunit•initiation factors is formed, i.e. that mRNA is bound after initiator-tRNA, or that a 30s subunit•mRNA•IF-3 complex is formed, which is then stabilized by the addition of formyl-methionyl-tRNAi and IF- I and IF-2. The next step in the process, the addition of the large subunit, does not require GTP, but
GTP hydrolysis is required to release IF-2 from the 70s ribosome complex. The role of GTP hydrolysis in initiation is still not fully understood, but it is not only required for the release of IF-2. Factor IF-3 is not only required for the binding of natural mRNAs, but also functions later either to dissociate the subunits after chain termination, or to keep dissociated subunits apart until another cycle of initiation is instituted. This activity of the IF-3 factor is referred to as its DF (ribosome dissociation factor) activity. None of the factors occurs on the polysomes and they recycle in protein synthesis, as shown in Fig. 10.6.
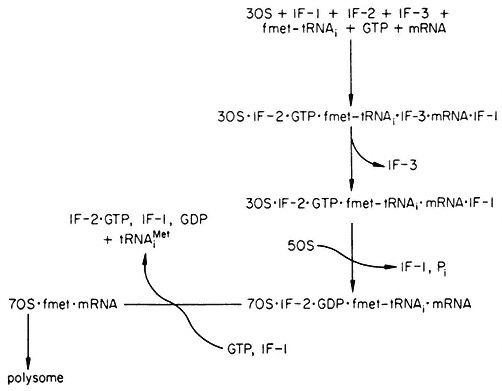
Figure 10.6
Recycling scheme for initiation factors in prokaryotes. The exact order
in which the factors interact with the 30s subunit is still not clear, nor is
whether fmet-tRNAi attaches before or after mRNA. IF-2•GTP•fmet-
tRNAi may exist independent of the 30s subunit. (fmet-tRNAi =acylated
formylmethionyl-tRNAi ; tRNAiMet =deacylated formylmethionyl-tRNAi ).
(Modified from Haselkorn & Rothman-Denes (1973).)
Several proteins which form an integral part of the small subunit have also been shown to be involved in initiation. Small subunit proteins, s1, s11, s13, s19, participate in IF-2 binding and s1, s11, s12, s13, s14, in IF-3 binding; s12 is responsible for recognition of mRNA (Haselkorn & Rothman-Denes, 1973; Anderson et al., 1974). Shine and Dalgarno (1974) have proposed a model of how the large subunit.initiation factor complex may recognize and attach mRNA. It would appear that the sequence (5') GGAGGU (3') is present in the same relative position with respect to the first translatable AUG triplet in all prokaryotic messengers so far analyzed. Furthermore, the 3' proximal end of 16s RNA has the sequence GAUCACCUCCU UA (OH), so that the underlined nucleotides could potentially base-pair with the GGAGGU sequence in the mRNA.
Further complexity of the initiation process is indicated by the work of
Schreier and Staehelin (1973), who have characterized five initiation factors (IF-E1 to IF-E5 ) for mammalian protein synthesis. Their results with rabbit globin mRNA are consistent with the formation initially of a methionyl-tRNA1 •IF-E2 •GTP complex, which is bound independently of mRNA to the 40s ribosome subunit by IF-E3 . Subsequent to this, mRNA and the 60s subunit are joined by the cooperative action of IF-E4 , ATP, IF-E1 and IF-E5 . The binding of natural mRNA requires IF-E4 and ATP. IF-E5 promotes the joining of the 40s complex with the 60s subunit and IF-E1 inhibits complex formation in the absence of mRNA binding. However, it has been suggested that the apparent need for additional factors results from deproteinization of ribosomal subunits and mRNAs, i.e. the requirement may be for structural proteins rather than initiation factors. The probable relationship between the different factors 1s given in Table 10.1. The little information available suggests that eukaryotic factors are not exchangeable with their prokaryotic counterparts.
| ||||||||||||||||||||||||||||||||||||||||
Factor EIF-3, contrary to its prokaryotic counterpart, consists of a number of different sized polypeptide chains; the question as to whether there are different EIF-3s or whether selectivity is controlled by additional mRNA specific factors, is still unresolved. A similar uncertainty to that described for the prokaryotic system surrounds the question as to whether or not methionyl-tRNAi binds to the 40s subunit prior to mRNA.
The initiation process is extremely complex and still not fully understood. A variety of proteins are involved, some being ribosomal structural proteins and others proteins which recycle during the process. These proteins interact, changing the conformation of the ribosome and thereby allowing the different steps of initiation to proceed in a sequential and orderly manner. Different mRNAs probably differ in their rate of attachment to the ribosome and/or in their efficiency in other stages of the initiation process, so affecting the frequency and rate of translation. With proteins which contain prosthetic groups their
absence may inhibit initiation, since it has been shown that lack of haemin in globin synthesis, results in the formation of an inhibitor of the binding of methionyl-tRNAi to 40s subunits (Adamson et al., 1972; Gross & Rabinovitz, 1973; Legon et al., 1973). Furthermore, a variety of mRNA specific factors have been proposed called 'interference' or 'i-factors', which affect the specificity of the initiation factor IF-3 (Groner et al., 1972). Thus, Strycharz et al. (1973) have identified a supernatant factor in Krebs II ascites cells, which is specifically required for the initiation of the translation of encephalomyocorditis viral RNA. However, Lodish (1974), from a kinetic analysis of protein synthesis, has proposed a model which precludes the necessity for mRNA specific factors. He points out that these have often been observed in systems which are less active than the corresponding in vivo system, and that the three eukaryotic systems available which translate exogenous mRNAs efficiently in vitro, do so with a variety of mRNAs.
10.3.2.2—
Elongation of the Polypeptide Chain
The elongation process starts once the 80s plant ribosome containing mRNA and methionyl-tRNAi , has been assembled and takes place by the following repeated cycle of events, each cycle being separated into:
(a) codon-directed binding of aminoacyl-tRNA;
(b) peptide bond formation; and
(c) translocation.
The ribosome has two binding sites; a 'P' site for the binding of methionyl-tRNAi and an 'A' site where all other aminoacyl-tRNAs bind. After the initiation complex has been formed, the next aminoacyl-tRNA, i.e. the one carrying the anticodon to the next codon of the mRNA, binds at the 'A' site. Once the aminoacyl-tRNA has been bound in the 'A' site, a peptide bond is formed by a peptidyl-transferase enzyme, which is part of the large subunit, such that the 'A' site now carries the aminoacyl-tRNA joined to methionine by a peptide bond. The 'P' site now carries the deacylated tRNAiMet , which is subsequently removed leaving an open 'P' site, and the peptidyl-tRNA moves from the 'A' to the 'P' site, so moving the message relative to the ribosome. The 'A' site is now empty and is ready for the next aminoacyl-tRNA to enter according to the next codon in the message. This cycle of reactions requires K+ , Mg2+ , GTP and various elongation factors, proteins, which are found in the soluble fraction of cell homogenates.
Our knowledge of polypeptide chain elongation is most complete with E. coli, where three elongation factors are known, EF-Tu, EF-Ts and EF-G (Fig. 10.7). Factor EF-Tu binds stepwise with GTP and aminoacyl-tRNA and this ternary complex transfers the aminoacyl-tRNA to the 'A' site of the ribosome, releasing an EF-Tu•GDP complex and inorganic phosphate. Factor EF-Ts displaces GDP to form the complex EF-Ts•EF-Tu, from which GTP displaces EF-Ts to regenerate EF-Tu•GTP. The third elongation factor, EF-G,
and free GTP, are responsible for the translocation of the peptidyl tRNA from the A' to the 'P' site with the prior removal of the deacylated tRNA from the 'P' site; GTP is hydrolyzed during the process. The elongation factors, like the initiation factors, recycle during protein synthesis (Fig, 10.7). The protein chain grows from the N-terminal end, and probably starts to fold into its three-dimensional conformation whilst the process of elongation is proceeding.
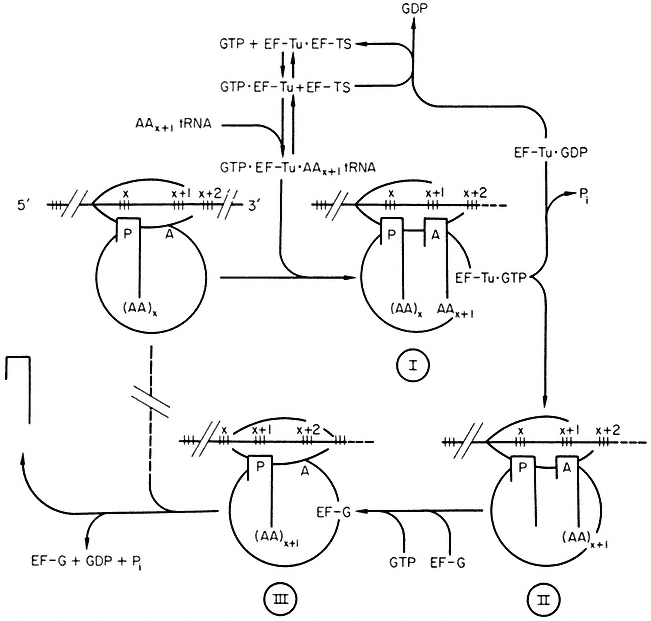
Figure 10.7
Polypeptide chain elongation in prokaryotes. Ribosomes cover more
than two codons on mRNA. Reaction I ® II is catalysed by peptidyl transferase.
(
(Modified from Haselkorn & Rothman-Denes (1973).)
Fewer details of the elongation factors are available in eukaryotic systems. However, two elongation factors, designated as the binding enzyme EF-1 and the translocase EF-2 (Hardesty et al., 1963; Gasior & Moldave, 1965) have been purified from ungerminated wheat embryos (Golinska & Legocki 1973; Twardowski & Legocki, 1973; Legocki, 1973; Allende et al., 1973). The mechanism of binding of aminoacyl-tRNA to wheat ribosomes is on the whole similar to
that in E. coli, the main difference being the absence in the eukaryotic system of a factor with the properties of EF-Ts, i.e. EF-1 = EF-Tu and EF-2 = EF-G.
Elongation factor EF-I, which requires the presence of Mg2+ at low concentration, K+ and GTP, is responsible for the binding of aminoacyl-tRNA to wheat ribosomes. It contains three active forms differing in molecular weight. The larger EF-1 forms dissociate to a single species of molecular weight 6 × 104 daltons during gel electrophoresis in the presence of SDS. The three forms observed may represent a monomer, trimer and tetramer of a single protein unit, of which the trimer seems to be the most stable form. Some experimental evidence supports the view that the light species is the active form of the EF-1 enzyme. For example, in calf brain and wheat embryo, it has been shown that the interaction of EF-1 with GTP and aminoacyl-tRNA yields a ternary complex which contains the light form of the enzyme, i.e. molecular weight 5 × 104 to 6 × 104 daltons (Moon et al., 1972; Legocki, 1973). It is suggested that the amino group of the esterified amino acid plays an important role in aminoacyl-tRNA binding (Jerez et al., 1969) and this differs from the bacterial system where EF-Tu can react with the deaminated product of phenylalanyl-tRNA, phenyl-lactyl-tRNA (Fahnestock et al., 1972).
The second elongation factor, EF-2, is involved in translocation of the peptidyl-tRNA from the 'A' site to the 'P' site during elongation. It is a protein of molecular weight 7 × 104 daltons and requires thiol compounds for activity (Twardowski & Legocki, 1973).
Ribosomes and elongation factors from the cytoplasm of eukaryotic organisms are not functionally interchangeable with their E. coli counterparts (Krisko et al., 1969). Furthermore, Perani et al., (1971) have shown absolute ribosome specificity, either for 70s or 80s ribosomes, of two sets of elongation factors, T and G (EF-1 and EF-2), isolated from yeast, and for factor EF-2 isolated from the alga, Prototheca zopfii. It is generally accepted that elongation factors within each ribosomal type (70s or 80s) are exchangeable between eukaryotes (Ciferri, 1972).
10.3.2.3—
Termination and Release of the Polypeptide Chain
No reliable information is available on chain termination in plants, most of the work having been done with E. coli. However, as there are some differences between the prokaryote and mammalian eukaryote systems, these will be mentioned briefly. In E. coli the elongation cycle is repeated until certain termination codons on the mRNA come into the 'A' site; these codons are UAG, UAA and UGA. At least three release factors, proteins, which recognize the terminator codons and bring about the release of the completed polypeptide chain from the tRNA, have been identified (Caskey et al., 1969; Capecchi & Klein, 1969; Milman et al., 1969). Originally designated R-1, R-2 and S these are now called RF-1, RF-2 and RF-3. RF-1 recognizes UGG or UAG, RF-2
recognizes UAA or UGA, and RF-3 affects the rate of release of the polypeptide chain. In mammalian systems, only a single release factor which responds to all three codons has been identified, and furthermore the prokaryotic termination requirement for GTP appears to be absent (Haselkorn & Rothman-Denes, 1973).
In addition, several other termination factors have been proposed from different organisms (see Haselkorn & Rothman-Denes, 1973). Although little is known about chain termination in plants, the presumed universality of the code suggests that similar terminator triplets and proteins are involved in plants also.
Originally, it was thought that on chain termination ribosomes dissociated into subunits and that the free ribosomes found in the cell were inactive in further protein synthesis. Noll et al. (1973) have presented evidence that at least some of the free ribosomes found in the cell interact with a factor and partially dissociate, so that the small subunit with bound IF-1, IF-2 and IF-3, can initiate protein synthesis.
In conclusion, it may be said that small differences exist between the mechanism of protein synthesis in the pro- and eukaryotic systems, e.g. the initiator tRNA, the involvement of GTP in different steps, as well as in the possible number of elongation factors. In view of the technical difficulties involved, work on the detailed mechanisms of eukaryotic systems could be considered hardly worthwhile. However, it is possible that translation level controls are much more important in eukaryotes than they are in prokaryotes. This possibility, together with the importance of plant proteins as a source of food, are justification enough for an attempt to characterize the process in detail in plants and, thereby develop suitable assay systems.
10.3.2.4—
Cell-Free Systems
In order to gain detailed knowledge of the mechanism of protein synthesis in plants, it is necessary to develop in vitro assay systems. Such systems must satisfy the following criteria:
(a) be dependent on the exogenous component(s) to be tested;
(b) be efficient, i.e. of comparable activity to in vivo rates, in order to allow quantitative analysis;
(c) the product, whose synthesis is monitored, must be clearly defined.
Two basic types of cell-free systems are used: fractionated systems, in which enzymes and components are isolated, purified and then reconstituted into an assay system, and unfractionated systems, in which the cells are broken open and cell-debris, nuclei and mitochondria removed by centrifugation leaving the ribosomes and the various components for protein synthesis in the supernatant, which is then used as the assay system. If all of the components of the assay are isolated from one organism, the system is said to be homologous as opposed to heterologous when they are not.
In addition to the fractionated E. coli prokaryotic system, there are four
fractionated eukaryotic systems which satisfy the above criteria. These are the rabbit reticulocyte (Lockard & Lingrel, 1969; Gilbert & Anderson, 1970), the mammalian liver (Prichard et al., 1971), the Krebs II ascites cell (Mathews & Korner, 1970), and the only plant system, that of wheatgerm (Allende & Bravo, 1966; Allende, 1970; Leis & Keller, 1970; Marcus et al., 1970a,b; Tarrago et al., 1970; Ghosh et al., 1971; Legocki & Marcus, 1970; Klein et al., 1972; Lundquist et al., 1972). Other fractionated cell-free systems from plants, such as those from developing legume seeds (Gumilevskaya et al., 1971; Payne et al., 1971a,b; Yarwood et al., 1971a,b; Beevers & Poulson, 1972; Wells & Beevers, 1973), are not well characterized and assays often lack quantitative accuracy. This is because in isolating the constituent enzymes and components, damage is caused in breaking the cell wall by hydrodynamic sheer, by cell vacuoles breaking and releasing acids, phenolics, tannins and other substances, and by activation of proteases and nucleases (Payne & Boulter, 1974). If the preparations used contain several components and structural damage to ribosomes has occurred, interpretation of the results may be qualitatively ambiguous, since enzyme preparations may contain proteins needed for the reconstitution of protein-leached ribosomes.
Unfractionated cell-free systems cannot be made from tissues which have a high nuclease or protease activity, and until better methods for the inhibition of degradative enzymes become available, most unfractionated systems from plants will be of limited value. Recently, Davies et al. (1972) have used media of high ionic strength and high pH with some success, and Gray and Kekwick (1973) have developed a system from pea seedlings using 0.2 mM vanadyl sulphate as an inhibitor. This system has been shown to synthesize the small subunit of ribulose bisphosphate carboxylase (Fraction 1 protein), since the tryptic peptides of the in vitro product were similar to those of the naturally occurring protein. The identity of the product was also proved by immunoprecipitation. The requirement of product identification is essential in complete cell-free assays (see also chapter 11, where the synthesis of the large subunit of Fraction I protein by the chloroplast cell-free system is described).
The wheatgerm system is by far the most successful unfractionated cell-free system from plants. Its preparation is as follows (Marcus et al., 1974):
Dry wheat embryos are ground thoroughly with a small amount of sand in a precooled mortar in a total volume of 3.3 ml of 90 mM KC1, 2 mM CaCl2 , 1 mM Mg (Ac)2 , 6 mM KHCO3 . The embryos are initially ground in 1.0 ml with 0.5 and 1.8 ml increments added subsequently. The slurry is then centrifuged for 10 minutes at 23,500 × g and the supernatant is removed with a Pasteur pipette, taking care to leave behind as much as possible of the upper lipid layer. Just prior to use, 0.5–3.0 ml are dialyzed against 500 ml of 1 mM Tris–acetate, pH 7.6, 50 mM KCl, 2 mM Mg(Ac)2 , 4 mM 2-mercapto-ethanol for 1.75 hours; this preparation is termed S23.
The wheatgerm system is attractive in spite of needing some additions, (e.g. tRNA), since it shows great promise as a 'translation' system for various mRNAs. This is an important development as the isolation of mRNAs from plants is now feasible using binding to oligo-dT columns (since a proportion of eukaryote messengers contain a poly-A sequence), by gradient centrifugation and by immunoprecipitation of polysomes (Haselkorn & Rothman-Denes, 1973; see also chapter 9).
Four RNAs of brome mosaic virus (BMW) induce amino acid incorporation into proteins when used as messengers in the wheatgerm system. RNA4 is translated with an efficiency comparable to that of bacteriophage RNA in E. coli extracts. The product is homogeneous and indistinguishable from the coat protein of BMV (Shih & Kaesberg, 1973). Satellite tobacco necrosis virus RNA has also been translated in this system and the product shown to be coat protein (Klein et al., 1972). Several natural eukaryotic messengers have also been translated in the wheatgerm system, e.g. rabbit globin mRNA (Efron & Marcus, 1973) and leghaemoglobin RNA (Verma et al., 1974). The latter is the only naturally occurring plant messenger to be isolated so far. It is a poly A-containing 9s–12s mRNA which was isolated from soybean root nodule polysomes. When used to programme the wheatgerm system by Verma et al., (1974), the product was identified serologically as mainly leghaemoglobin S (LbS ) with a little leghaemoglobin F (LbF ); there are two types of leghaemoglobin found in soybean nodules and the reason for the preferential synthesis of the LbSin vitro, is not understood. Factors other than those already present in the wheatgerm system were not required, and there was no need for the addition of heme.
There are several unfractionated cell-free systems from animals (Haselkorn & Rothman-Denes, 1973). Of particular interest is that of the Xenopus oocyte since intact eggs are used to translate exogenous messages (Gurdon et al., 1971). Natural plant messages have not been used but there is evidence that tobacco mosaic virus RNA can be translated (Knowland, 1973). Coat protein is not synthesized, however, and the main product is a polypeptide with molecular weight 14 × 104 daltons. Although no function has been assigned to this protein, it is known that plant cells infected with TMV also make a protein of the same molecular weight (J. Gurdon, personal communication).
The use of heterologous systems assumes the interchangeability of the various enzymes and components between different systems and as mentioned previously, it is generally accepted that this is not usually possible between prokaryotic and eukaryotic organisms. Generally, attempts to demonstrate interchangeability between different eukaryotic systems have been positive, but several workers have suggested that this is because discrimination can only be seen under optimum conditions. The general conclusion is that tissue and organism specificity may occur but only in exceptional cases (Mathews, 1973).
10.3.2.5—
Inhibitors
Several antibiotics and other inhibitors block various steps in protein synthesis, and are, therefore, extremely useful in establishing the mechanism of the process (Pestka, 1974). Table 10.2 lists the more common inhibitors and their sites of action; many of these have been used with plants. It is important to realize
| ||||||||||||
that the concentration of the inhibitor is often critical for the production of a specific effect; it is essential to establish the conditions for the correct use of an inhibitor with each tissue and to maintain effective controls. It has been suggested by Glazer and Sartorelli (1972) that in rat liver, membrane-bound 80s ribosomes are more susceptible to a range of inhibitors than are the 80s ribosomes free in the cytosol.