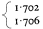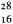Chapter 9—
The Nucleus and the Organization and Transcription of Nuclear DNA
9.1—
Introduction
The importance of the cell nucleus has been recognized for some considerable time and a great deal of literature concerned with its general organization and composition and the structure and behaviour of chromosomes has been derived from cytological investigations. This present chapter is mainly concerned with recent developments in research which provide a more detailed understanding of the organization and function of nuclear DNA, the mechanism of RNA synthesis and the question of the control of gene expression in higher plants.
9.2—
Nuclear Structure and Composition
In general, nuclei are spherical or disc-shaped, although occasionally they may appear lobed. They vary in size in different species but are usually within the range 1–10 µm in diameter (Fig. 9.1). Nuclei may be isolated by gently breaking cells in a suitable osmoticum, filtering the homogenate, for example through muslin, followed by low-speed centrifugation. Attempts are usually made to free the nuclei of contaminating cytoplasm, cell wall fragments and other organelles by centrifugation through sucrose. However, methods for obtaining high yields of intact, purified nuclei from higher plant cells are at present not very satisfactory. For certain purposes it is often easier to work with chromatin preparations which consist essentially of chromosomal and nuclear material obtained from broken nuclei.
The major part of the cellular DNA is present in the nucleus, arranged in a highly organized fashion within the chromosomes which at different times may show characteristic bands or loops (see Fig. 9.1). Chromosomes are composed of approximately equal amounts of DNA and basic proteins known as histones (see Table 9.7). In addition, much smaller quantities of RNA and acidic proteins are present which are probably of great importance in controlling gene expression. There is wide variation in the number of chromosomes observed within the nuclei of different species although in diploid cells of a single species the number is normally constant. During interphase the chromosomes are less highly condensed than at mitosis and are more difficult to detect. At this stage at least part of the DNA is being transcribed into RNA. It is possible to identify chromosomes on the basis of morphological features such as size, position of the centromere and the presence or absence of constrictions. In addition, certain
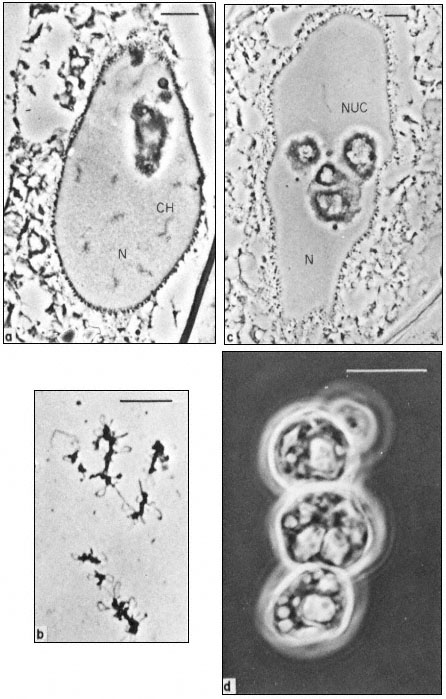
Figure 9.1
Phase contrast light micrograph of nuclei and isolated nuclear
contents from Acetabularia. These giant nuclei are unusually large;
the scale line represents 10 µm. Chromosomes (CH) and nucleoli (NUC)
can be seen within the nuclei (N) fixed in situ (a and c) and the appearance
of isolated chromosomes and nucleoli are shown in b and d. Note the large
aggregates of nucleolar material with an internal cavern system (c and d) and the
'lampbrush-chromosome-like' morphology of the chromosomes (b). (Fig. 9.1a, c and d
from Spring et al., 1974; 9.1b, previously unpublished micrograph from the same authors.)
areas of heterochromatin, which appear more highly condensed and therefore take up more stain, may be distinguished from the remainder of the chromosome, termed euchromatin. In certain cell types, such as embryo suspensor cells, giant polytene chromosomes occur. In such situations it is possible to study the structural organisation of chromosomes in much greater detail.
Recent developments have made it possible to identify individually each of the 46 chromosomes in man on the basis of characteristic banding patterns produced by certain staining procedures. These techniques and their application to the study of plant chromosomes have been reviewed by Vosa (1975).
Organelles known as nucleoli become visible within the nucleus during interphase. These are very prominent spherical bodies with different staining properties to the remaining part of the nucleus. They arise in association with specific regions, the nucleolar organizers, of certain chromosomes. Nucleoli are not surrounded by a membrane but they do have a highly ordered structure (Fig. 9.2). The central core consists of a region of fibrils 8–10 nm across surrounded by a region of dense granules 15–20 nm in diameter. Mutants which lack nucleoli are unable to synthesise ribosomes and it is now known that the genes coding for ribosomal RNA normally occur at the nucleolar organizer region of chromosomes. Here the DNA is transcribed into ribosomal-RNA (rRNA) which is assembled to form ribosomes in the outer region of the nucleolus by combination with ribosomal proteins. The nucleolus is most prominent, therefore, when synthesising large numbers of ribosomes. This explains the old observation that there is a general correlation between nucleolar volume and the rate of protein synthesis of cells.
The fact that nuclei are surrounded by a membrane was deduced from the observation that they are selectively permeable to small molecules. The nature of the double membrane may be readily observed with the aid of the electron microscope (Roberts & Northcote, 1971). It is often interconnected with the cytoplasmic membrane system, the endoplasmic reticulum, and is perforated by a large number of pores. This is demonstrated quite clearly in Fig. 9.3 which shows a surface view of a portion of the nuclear membrane of Acetabularia. The apparent diameter of a nuclear pore is of the order of 100 nm. However, the observation that colloidal particles of much smaller size do not readily pass through these pores suggests that their effective size is substantially less, probably due to the presence of a protein-containing matrix within the pore.
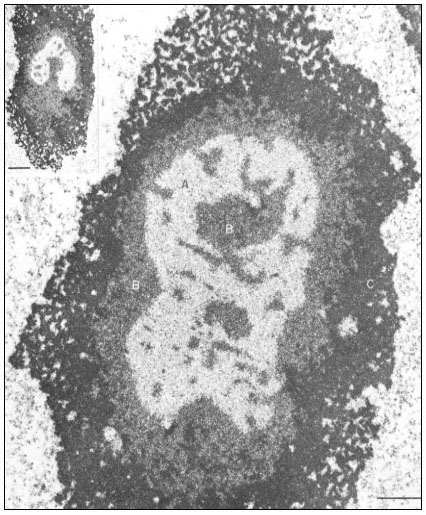
Figure 9.2
Ultrastructural details of the composition of the nucleolar subunits of Acetabularia
revealed by electron microscopy. Note the finely fibrillar texture in the internal zone
A and the dense packing of small granulofibrillar structures in zone B, extensions
from which deeply penetrate zone A (see also the insert ) and the very dense
packing of granules in the cortical zone C. The scale indicates 1 m m,
the scale in the insert denotes 2.5 µm. (From Spring et al., 1974.)
9.3—
Nuclear DNA Content
The amount of DNA present in nuclei may be estimated by nuclear isolation followed by extraction and direct chemical measurement of DNA content. A more elegant approach is to carry out quantitative microspectrophotometry of
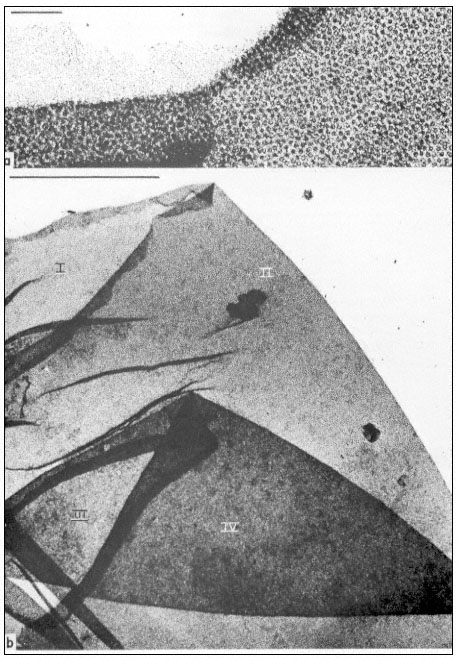
Figure 9.3
Manually isolated nuclear envelope of Acetabularia mediterranea. This shows
a sequence of overfoldings (I to IV in b). Note the high density of nuclear pore
complexes and the association with some fibrillar material as identified in the
upper region of a. The scale in (a) denotes 1 µm, the scale in (b) denotes 10 µ m.
(From Spring et al., 1974.)
individual nuclei following Feulgen staining. Table 9.1 compares the nuclear DNA content of a range of higher plants with the amounts of DNA found in lower organisms. Two conclusions can be drawn from such a comparison. Firstly, and perhaps not surprisingly, higher plants contain considerably more DNA than bacteria and fungi. For example, Pisum sativum contains 1,000 times as much DNA as Escherischia coli . Secondly, however, the amount of DNA present in different higher plants may vary by as much as 100 fold. Further comparison shows that there is little correlation between chromosome number and DNA content.
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ||||||||||||||||||||||||||||
One explanation of the increased DNA content of higher plants is that more complex organisms require a greater store of genetic material than bacteria and viruses. Although it is difficult to estimate precisely the extent of the increased coding requirement it is quite clear that this is in itself not a satisfactory explanation for the presence of such large amounts of DNA in some plants. There often exists a wide variation in the 2 C DNA content of plant species within one: genus and it is unreasonable to conclude that small differences between related plants are a direct consequence of gross differences in nuclear DNA. Furthermore, some plants contain 20 times as much DNA as mammals and yet the latter are normally accepted as comprising a more complex group of organisms. It therefore appears that the amount of nuclear DNA is not directly related to the complexity of the organism. It is also possible to argue on other grounds that the total amount of nuclear DNA is not required in order to code for additional proteins. Firstly, it is difficult to envisage functions to account for a 1,000 fold increase in the number of enzymes. Secondly, if each higher plant contained such a large number of essential genes it would, on numerical grounds, be prone to an unrealistically high incidence of mutation.
The size of the genome of plants with relatively low nuclear DNA contents is of the order of 2 × 1012 daltons. Given that the molecular weight of a gene required to code for a messenger-RNA (mRNA) molecule for an average-sized polypeptide is 106 daltons, this represents, in principle, 2 × 106 genes. A more realistic estimate would be that 1% of this DNA actually codes for proteins. Experimental evidence for this view and a discussion of the organization, properties and possible function of the remaining DNA is given later.
It used to be widely proclaimed that when differences in ploidy are taken into account the amount of DNA in each nucleus of a single species is constant. The observation that the gene number remains constant, and the fact that many plant cells are totipotent, supports the view that differentiation involves the selective control of gene expression. However, the rule of DNA constancy is not inviolate. It is now known that specific regions of the genome may become selectively amplified at certain stages of development. This undoubtedly occurs to the genes for ribosomal RNA during oocyte development in Amphibia. Although there exist no established cases of ribosomal RNA gene amplification in higher plants, the satellite DNA present in melon may be under- or over-replicated in different tissues (Ingle & Timmis, 1975).
One very intriguing observation is that the amount of nuclear DNA in flax may become altered if the plants are subjected to certain regimes of nitrogen and phosphorus fertilizer. The alteration in nuclear DNA content is stable and inherited (Evans, 1968). Although this phenomenon is well established it is not known how widespread it may be and there is no satisfactory explanation to account for it.
9.4—
Fractionation and Properties of DNA
9.4.1—
General Properties
Although single-stranded DNA does occur naturally most DNA molecules consist of two anti-parallel chains of nucleotides. Each nucleotide consists of deoxyribose, a phosphate group and a base. The four bases that occur in all types of DNA are the purines adenine and guanine and the pyrimidines thymine and cytosine. Adenine is capable of forming hydrogen bonds with thymine, and guanine with cytosine. Sometimes modified bases are present in small amounts and in plant nuclear DNA a substantial proportion of the cytosine residues are replaced by 5-methyl-cytosine. In double stranded DNA there is an equivalence between the amount of adenine and thymine and guanine and cytosine plus 5-methyl-cytosine. The two strands are joined together by hydrogen bonds between complementary pairs of bases to form a right handed helix. The bases are at right angles to the sugar residues, and project inwards, and the phosphate groups are on the outside. In the B form of DNA there are ten base pairs in one complete turn of the helix, which extend for 3.4 nm. The width of the helix is 1.8 nm. An alternative configuration is possible, the A form, where the bases are tilted and 11 pairs are accommodated per turn of the helix. Double stranded RNA and DNA-RNA hybrids adopt this latter configuration.
Many DNA molecules are known to be several millimetres in length. It is very difficult to determine the actual length of very large DNA molecules because they are extremely sensitive to shearing forces produced by various isolation procedures. It is known that the complete genome of E. coli, with a molecular
weight of 2.9 × 109 daltons, consists of a single circular DNA molecule of length approximately 1 mM . The total amount of DNA in a higher plant cell is equivalent to a length of approximately 1 m, but it is not known how many molecules this comprises.
The base composition of DNA is normally expressed as the per cent G + C content. This value, which is characteristic of a given species, varies widely in different organisms and in bacteria may be anything from 30% to 70%. The base compositions of a number of plant DNAs are given in Table 9.2.
9.4.2—
Buoyant Density Centrifugation
One of the most common methods of fractionating DNA is to exploit differences in density between different types of DNA in concentrated salt solutions. Caesium chloride is the most commonly used. The method involves prolonged high speed centrifugation of purified DNA in concentrated caesium chloride. The centrifugation produces a gradient of salt concentration in the centrifuge tube and the DNA migrates to occupy a position in the gradient which corresponds to its own buoyant density. A number of factors affect the buoyant density, such as the nature of the caesium salt, the presence of heavy metals or DNA-binding dyes, the pH and the temperature. Under constant conditions (usually 25°C in caesium chloride at neutral pH) the buoyant density of DNA is related to the GC content:
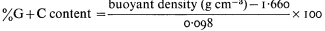
DNAs with different base compositions can therefore be separated by this method. Figure 9.4 shows the fractionation of Phaseolus aureus DNA (buoyant density = 1.695) and E. coli DNA (buoyant density = 1.710) in caesium chloride.
Most nuclear DNAs from higher plants have buoyant densities within the range 1.69–1.71 g cm–3 . However, the presence of 5-methyl-cytosine serves to reduce the density slightly, thereby giving rise to an under-estimate of the GC content. In general, 1% methylation decreases the buoyant density by 1 mg cm–3 . Certain sequences of bases may also distort the relationship between base composition and buoyant density. Furthermore, single-stranded DNA is denser than double-stranded DNA of similar base composition by approximately 0.015 g cm–3 and under alkaline conditions the density is increased by 0.06 g cm–3 . This is due partly to the fact that DNA becomes single-stranded under these conditions and also partly because the deprotonated adenine and thymine residues are neutralized by binding caesium ions.
9.4.3—
Satellite DNA
Figure 9.4 shows that DNA from a particular organism often forms an apparently homogenous band in caesium chloride. The width of the band depends
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
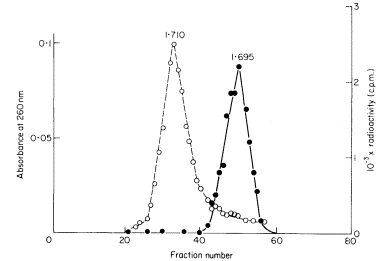
Figure 9.4
The separation of a mixture of E. coli and P. aureus DNA in caesium chloride.
A trace of E. coli DNA labelled with 3 H thymidine was mixed with P. aureus DNA
and centrifuged at 30,000 rpm at 25ºC in caesium chloride for 3.5 d using a fixed-angle
centrifuge rotor. Three-drop fractions were collected and the P. aureus DNA located
by absorbance at 260 nm (solid circles, continuous curve) and E. coli DNA detected by
measurement of radioactivity (open circles, dotted curve). (Unpublished result of D. Grierson.)
upon the slope of the density gradient, the molecular weight of the DNA and the difference in base composition between different regions of the DNA. In most plants certain regions of the genome are sufficiently different in base composition for them to occupy a position distinct from the main-band DNA. If they represent only a small percentage of the total DNA such sequences are difficult to detect but when present in large amounts they produce one or more peaks of satellite DNA. Melon DNA provides an example where the satellite is denser than the main-band (Fig. 9.5) but light satellites have also been observed, for example in flax. It has been assumed that the majority of satellite DNAs are nuclear in origin but recent studies with cucumber suggest that they may sometimes represent organelle DNA (Kadouri et al., 1975). Although they are widely distributed in plants no satellites have so far been found in monocotyledonous plants using caesium chloride fractionation (Table 9.3). However similar components can be detected using other techniques. For example, substances such as heavy metals and dyes often selectively bind to certain sequences within the DNA. This alters their buoyant density and thus generates satellites. One common method is to fractionate DNA in caesium sulphate gradients in the presence of silver ions. Using this procedure, Ingle (unpublished) has demonstrated the presence of a satellite DNA in Scilla (a monocotyledon) although in caesium chloride no such satellite is observed.
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
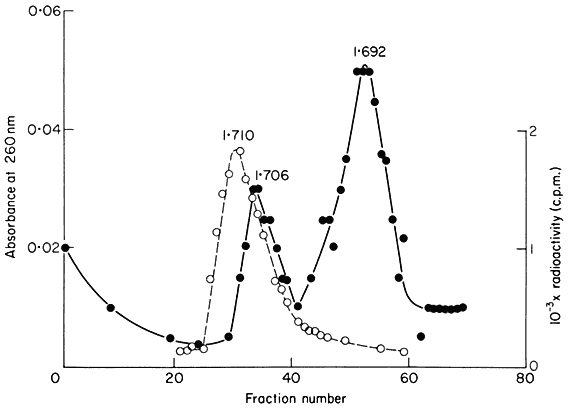
Figure 9.5
The separation of a mixture of E. coli and melon DNA in caesium chloride.
The experimental details were as described in the legend to Fig. 9.4. Melon
DNA was detected by absorbance at 260 nm (solid circles, continuous curve)
and E. coli DNA by measurement of radioactivity (open circles, dotted curve).
(Unpublished result of D. Grierson.)
9.4.4—
Dissociation and Reassociation of DNA
When double-stranded DNA in solution is heated above a certain temperature the hydrogen bonds between complementary base pairs are broken and the two strands dissociate. The process, which can also be brought about by treatment of DNA with alkali, is commonly referred to as melting or denaturation. Strand separation is accompanied by an increase of approximately 35% in the absorbance of ultra-violet light. This hyperchromic effect provides a means of monitoring the dissociation of double-stranded DNA by measuring the increase in absorbance of a DNA solution at 260 nm as the temperature is raised. The melting curves of DNA from a variety of sources are compared in Fig. 9.6. The temperature at which 50% of the DNA is melted, the Tm, depends upon a number of factors such as the salt concentration and pH. Under standard conditions, i.e. when the DNA is dissolved in standard saline citrate (S.S.C. being 0.15 M sodium chloride, 0.015 M trisodium citrate, pH 7) the base composition and the Tm are related in the following way:
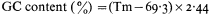
In viruses, the majority of sequences within the genome are similar in base composition and for a given DNA a sharp melting profile is observed with a characteristic Tm. In contrast, higher organism DNA often consists of distinct subfractions, each with a slightly different Tm. This produces a broad melting
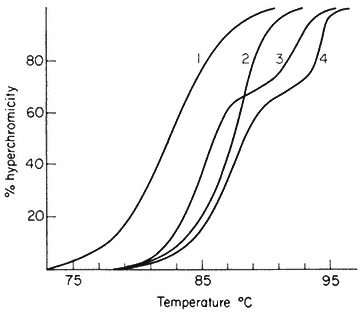
Figure 9.6
Melting curves of various DNA samples. (1) Musk melon main-
band and (4) satellite DNA. (3) a 2:1 mixture of DNA from bacterium
C and phage P58. (2) B. subtilis DNA. Melting was carried out in S.S.C.
(From Bendich & Anderson, 1974.)
profile which represents a number of overlapping melting curves. With DNA samples from organisms that contain prominent satellite bands in caesium chloride the melting curve often has two or more phases. Even where no large satellites are present heterogeneity within the DNA can readily be detected by plotting the rate of change of absorbance at 260 nm of a DNA solution as a function of temperature.
If, following denaturation of DNA by alkali or by heating, the pH of the solution is restored to neutrality or the temperature reduced below the Tm, complementary strands of DNA show a marked tendency to recombine to form double-stranded molecules. This process of duplex formation is termed renaturation or reassociation. The rate of the reaction is governed by the concentration of DNA sequences, pH, temperature and salt concentration. Renaturation is normally carried out at moderately high salt concentrations at a pH close to neutrality and at a temperature approximately 25°C below the Tm. Under these conditions, single-stranded DNA molecules collide at random. Most collisions occur in such a way that complementary sequences of bases are not in register and the strands do not reassociate. If, however, two strands approach in an anti-parallel configuration and potential regions of complementarity are in close proximity a nucleation event will take place and hydrogen bonds will form. Whether or not the strands remain together depends upon the nature of the neighbouring bases on either side of the nucleation site. If complementary sequences are arranged in the correct register along the length of the molecule a rapid zippering effect takes place to produce a stable double-stranded DNA molecule.
DNA renaturation can be followed by measuring any property that distinguishes single-stranded from double-stranded DNA. The decrease in absorbance at 260 nm that accompanies duplex formation is often exploited for
this purpose but fractionation by hydroxyapatite is also commonly used. Under appropriate conditions only double-stranded DNA binds to hydroxyapatite (a hydrated form of calcium phosphate); single strands pass straight through a column and double-stranded material may subsequently be eluted either by increasing the salt concentration or by raising the temperature above the Tm. This procedure has the advantage that the properties of the single-stranded and renatured fractions may subsequently be studied separately.
9.4.5—
Repeated Sequence DNA
Information about the sequence content of DNA can be obtained by studying the renaturation rate. Double-stranded DNA is first sheared to lengths of a few hundred nucleotides by ultrasonication or by forcing through a narrow gauge needle. This is designed to liberate short sequences of DNA and to allow them subsequently to react independently of neighbouring sequences. The DNA is then denatured and allowed to renature at the desired concentration under controlled conditions. Renaturation is followed by monitoring the absorbance of the solution at 260 nm in a spectrophotometer cell maintained at the appropriate temperature, or alternatively by taking samples at intervals and fractionating them by hydroxyapatite chromatography. In some situations partially double-stranded segments of DNA are produced, which have single-stranded 'tails'. These can be removed by treating the reassociated DNA with SI nuclease, which is specific for single-stranded DNA, before hydroxyapatite fractionation.
The reassociation reaction follows second order kinetics, which means that it is governed by the concentration of the reacting sequences. This is determined by the DNA concentration and the sequence content. For example, synthetic poly-dA. dT renatures almost instantaneously, whereas at the sme concentration naturally occurring DNA takes much longer to renature. In general the renaturation rate is inversely proportional to the number of base pairs in the genome (providing that no repetitive sequences are present). In principle, therefore, the number of 'genes' or sequences may be estimated by comparing the renaturation rate of an unknown DNA with that from an organism with a well characterized genome. The results of renaturation experiments are often expressed as a 'Cot' plot. In this form the extent of DNA renaturation is plotted as a function of the initial concentration of the DNA (in moles of nucleotides per litre) multiplied by the renaturation time (in seconds) plotted on a log scale. One advantage of this is that different types of DNA can easily be compared by their Cot 1/2 values (the value of Cot at which 50% of the DNA becomes renatured) (Fig. 9.7a). For a variety of microorganisms and viruses the Cot 1/2 is related to the genome size. However, this is not strictly true for eukaryotic DNA which behaves in a more complex way.
In bacteria and viruses most DNA sequences are represented only once in the genome but in higher organisms many sequences occur in multiple copies. For this reason a plot of the renaturation of higher plant DNA deviates markedly
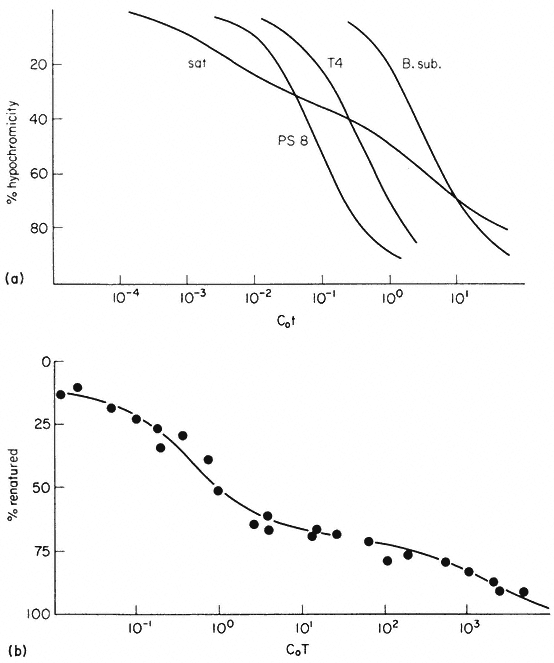
Figure 9.7
Reassociation kinetics of various DNA samples. (a) Musk melon satellite
(sat.), phage PS8 (PS8), Bacillus subtilis (B. sub. ), phage T4 (T4). The data
are corrected for differences in GC content and to take account of different
solvent concentrations. (b) Helianthus annuus DNA. The slowest reassociating
fraction behaves approximately as expected for sequences present in one copy per
haploid genome. (a) redrawn from Bendich and Anderson, 1974; (b) from Flavell et al., 1974.)
from a second order curve (Fig. 9.7b). A substantial proportion of the genome is so highly repetitive that it renatures even more rapidly than bacterial DNA. A second, intermediate, fraction contains families of repetitive sequences present in a lower frequency. Finally, there exists a unique fraction of DNA which essentially comprises sequences present in only one or a small number of copies in the genome. Flavell et al., (1974) have studied the reassociation kinetics of the DNA from 23 higher plant species, which vary in 2C nuclear DNA content between 1.7 and 98 × 10–12 g. The results, summarized in Table 9.4, show that in plants with relatively large genomes an average of 80% of the DNA consists of sequences present in 100 copies or more. For plants with smaller genomes
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
the average amount of repetitve DNA is 62%. These results indicate that part of the variation in nuclear DNA mass can be accounted for by variation in the amount of repeated sequence DNA.
9.5—
The Genes for Ribosomal-RNA
More is known about the properties, organization and function of the genes for rRNA than any other part of the genome. These DNA segments direct the synthesis of rRNA molecules which form the structural 'backbone' of ribosomes, without which cells cannot function. It has been known for some time that they are associated with the nucleolus because certain mutants of the toad Xenopus, which lack nucleoli, are incapable of synthesizing rRNA. During development these mutants die because they are unable to synthezise ribosomes.
9.5.1—
DNA-RNA Hybridization
The genetic origin of any RNA molecule can be studied by DNA-RNA hybridization. This technique is similar in principle to DNA-DNA reassociation. It depends upon the fact that any RNA molecule synthesized on a DNA template has a base sequence complementary to the DNA strand from which it was transcribed. Under suitable conditions, therefore, DNA can be induced to combine with complementary-RNA (cRNA) to produce an RNA-DNA double helix. This provides a means of locating and measuring the genes that code for a particular type of RNA.
One of the commonest procedures is to prepare pure DNA, denature it by heating or treatment with alkali, and to attach the single strands to a solid matrix such as a nitrocellulose filter. During subsequent treatments the immobilized DNA strands are prevented from reassociating but they are capable of forming hybrids when they are placed in a solution of nucleic acid containing complementary base sequences. Using radioactive RNA the formation of DNA-RNA hybrids can be detected by measuring the amount of radioactivity that becomes associated with the filter. The reaction is affected by a number of factors such as RNA concentration, salt concentration, temperature, the presence of compounds such as formamide, which lowers the Tm, and the reaction time. With rRNA at a concentration of 2 µg cm–3 at 70°C in 6 × S.S.C. the reaction is virtually complete within a couple of hours (Fig. 9.8). After a suitable time the filters are washed, given a mild treatment with ribonuclease to remove any unhybridized RNA, and the amount of radioactive RNA remaining is measured in a liquid scintillation counter. If the amount of DNA initially applied to the filter is known then the extent of hybridization can be calculated from the specific radioactivity of the RNA. The amount of DNA that hybridizes to rRNA can either be calculated directly from the saturation plateau (Fig. 9.8) or, more accurately, from a double reciprocal plot of the data (Scott & Ingle, 1973). In most plants between 0.05 and 1% of the DNA hybridizes to rRNA. This represents from a few hundred to several thousand genes per diploid nucleus, depending on the species (Table 9.5). The 258 and 18S rRNA molecules hybridize to separate sites on the DNA in accordance with the ratio of their molecular weights (1.3:0.7; approximately 2:1 by mass but 1:1 molar ratio).
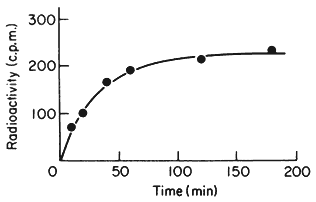
Figure 9.8
Saturation hybridization of nuclear DNA from Phaseolus
aureus with radioactive 25S rRNA. 25s rRNA (molecular weight
1.3 × 106 daltons) labelled with [3 H]-uridine at a specific activity
of 1.4 × 104 c.p.m. µg–1 was hybridized to nuclear DNA (2 µg
per filter) for increasing periods of time in 6 × S.S.C. at 70°C.
(From Grierson, 1975).
| |||||||||||||||||||||||||||||||||||||||||||||||||||
This indicates that there are equal numbers of genes for each type of rRNA. In cases where they do not hybridize in equimolar amounts this may be due to sequence homology between the two RNAs, or alternatively may arise by contamination of the RNA samples with other types of RNA.
Multiple genes for rRNA are a physiological necessity in order to provide sufficient sites of synthesis for rRNA molecules to meet the high metabolic demand for ribosomes. Plants often contain thousands of these genes although the number varies between varieties of the same species and between different species. Most of the available evidence suggests that all the copies are identical and this provides a clear example of repetitive DNA with a known function.
Their localization within the nucleolus has been confirmed by in situ hybridization of rRNA to cytological preparations (Brady & Clutter, 1972).
In many eukaryotes ribosomal DNA occupies a separate position in caesium chloride gradients relative to main band DNA. However, since they represent 1%, or less, of the genome these genes are not easily detected by studying the distribution of the DNA. Nevertheless, they can readily be pinpointed by hybridizing rRNA with separate fractions of DNA from the gradient. The fractionation of main band DNA from the rRNA genes of Phaseolus aureus is shown in Fig. 9.9. In general, ribosomal DNA from plants has a higher density
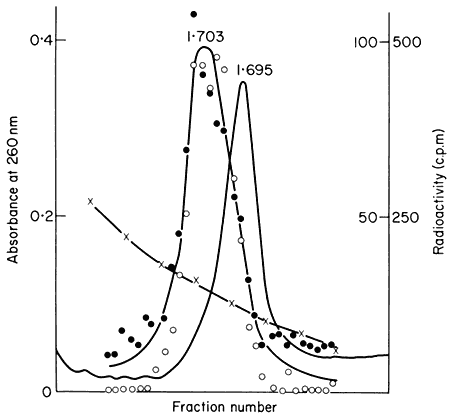
Figuer 9.9.
Separation and detection of the DNA sequences coding for
ribosomal RNA. Nuclear DNA from Phaseolus aureus was
fractionated in caesium chloride (see Figs. 9.4 and 9.5) and
the fractions from the gradient were divided into two and
hybridized to either 25s (cytoplasmic) or 23s (chloroplast)
rRNA labelled with [3 H]-uridine. Smooth, curve, absorbance
of DNA at 260 nm; solid circles, hybridization with 25s rRNA;
open circles, hybridization with 23s rRNA. The gradient of CsCl
is shown and the calculated densities of the rRNA genes and
the main band DNA are indicated. (Unpublished result of D. Grierson.)
than main band DNA. This is consistent with the fact that in higher plants rRNA has a base composition in excess of 50% GC, whereas the bulk of nuclear DNA often has a lower GC content. In this sense rRNA genes represent a type of satellite DNA. However, in different plants the rRNA genes are often present in similar numbers and with a similar density irrespective of the presence or absence of large amounts of satellite DNA. It therefore appears that ribosomal DNA and other satellites may occasionally band together in caesium chloride but this is entirely fortuitous (Ingle et al., 1975).
9.5.2—
Electron Microscopy
The rRNA genes can also be studied by electron microscopy of preparations from isolated nucleoli. This approach provides a clear indication of their organization. Figure 9.10 shows an electron micrograph of the partially
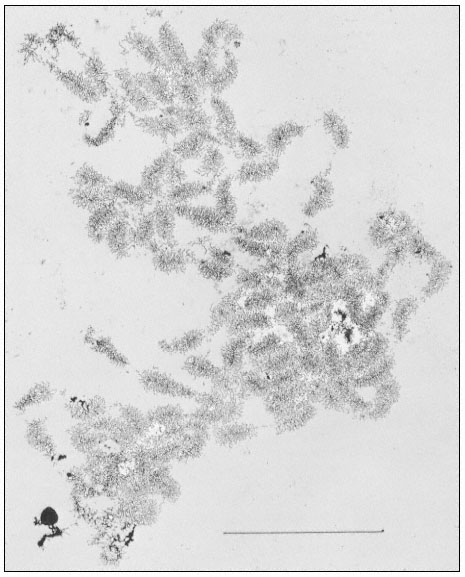
Figure 9.10
Spread contents of a complete nucleolar aggregate subunit from Acetabularia
mediterranea. Note the numerous 'matrix' units. The scale indicates 5 µm.
(From Spring et al., 1974.)
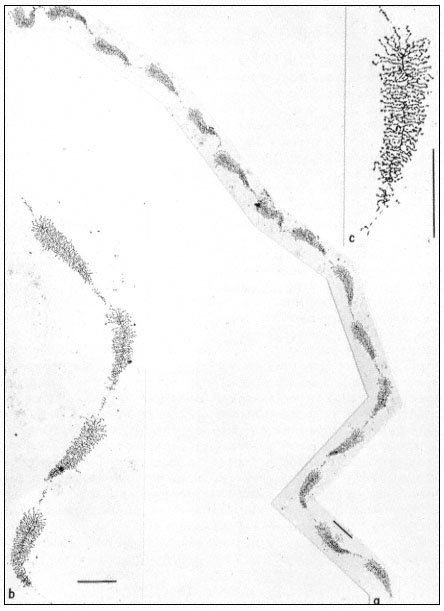
Figure 9.11
(a) Well spread nucleolar material showing the regular pattern of 'matrix'
units and 'spacer' segments. Note the length gradient of lateral fibrils within
each matrix unit. (b) Note occasional groups of small fibrils associated with the
spacer units. (c) At higher magnification, the dense packing of lateral fibrils within
the matrix unit is clearly seen. Scales indicate 1 µm, (From Trendelenburg et al., 1974.)
dispersed contents of a subunit from the nucleolar aggregate of Acetabularia. This shows a regular arrangement of fibril-covered 'matrix' units and fibril-free 'spacer' regions arranged along a central axis. A higher magnification of a 'matrix' unit is shown in Fig. 9.11. The central axis is the double-stranded ribosomal DNA itself. The 'matrix' units represent ribosomal genes which are being transcribed into RNA at a number of points simultaneously. The fibrils of the 'matrix' unit are growing chains of rRNA precursor molecules (see the section on RNA synthesis). RNA polymerase enzymes can be seen clustered on the DNA. About 120 enzymes are engaged in the simultaneous transcription of the DNA in each 'matrix' unit. The marked polarity reflects the fact the growing chains of attached RNA molecules increase in length as the polymerase enzymes responsible for their synthesis proceed along the DNA axis. In addition there is a tendency for the free ends of the RNA chains to form terminal knobs as they become folded, probably in association with proteins.
Multiple rRNA genes therefore alternate in a precise arrangement with non-transcribed regions of the DNA. Hybridization evidence shows that separate 'cistrons' for the 25s and 18s rRNA are present in the DNA, but there is no evidence for the existence of separate transcription units. In contrast, in Acetabularia each transcribed region of double-stranded DNA is approximately 2 µm long. This indicates that it codes for an RNA molecule with a molecular weight of approximately 2 × 106 daltons. This is the combined mass of the 25s and 18s rRNA molecules (1.3 + 0.7 × 106 daltons). There is clear evidence (discussed in the section on RNA synthesis) that ribosomal RNA is first transcribed as a polycistronic precursor molecule. In many plants this precursor has a molecular weight of approximately 2.4 × 106 daltons, and contains one sequence of 25s RNA plus one sequence of 18s RNA together with some extra sequences which are subsequently removed. In order to reconcile such a mode of synthesis with studies on the organization of the rRNA genes it is necessary to propose that each 'matrix' unit consists of an 18s and 25s sequence and some transcribed 'spacer' DNA. Fortunately, in cases where the organization of ribosomal DNA has been studied in detail, for example in Xenopus, evidence for precisely this type of arrangement has been obtained (Birnstiel et al., 1968). Furthermore, recent studies with the electron microscope have provided evidence for a common organization of the rRNA genes in all animals examined (Schibler et al., 1975). A model for the repeating unit, which probably applies also to plants, for it is compatible with all the available evidence, is shown in Fig. 9.12.
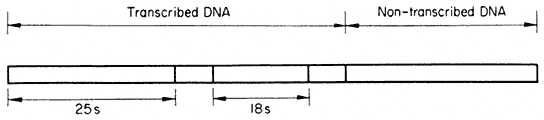
Figure 9.12
Suggested structure of the repeat unit of ribosomal DNA in plants.
(Modified from Schibler et al., 1975.)
By carrying out partial denaturation studies it has proved possible to map certain areas rich in GC or AT base sequences. In addition, it is known that both the non-transcribed 'spacer' regions and the transcribed spacer can vary significantly in length and composition between species. Possible functions for such regions are discussed below. In contrast, however, the regions that code for rRNA are remarkably constant in composition. This probably reflects the importance of their role in providing structural RNA for ribosome formation.
As shown in Fig. 9.9, chloroplast rRNA also hybridizes to nuclear DNA (Ingle et al., 1970; Tewari & Wildman, 1968). This is a surprising result in view of the fact that chloroplasts contain their own DNA and synthesize their own rRNA (see chapter 11). The location of these genes within the nucleus is not known, and at present there is no evidence that they are ever transcribed in the nucleus.
9.6—
Organization and Function of the Genome
9.6.1—
Repetitive DNA
Reassociation studies indicate that between 40% and 90% of the genome of higher plants is composed of repetitive DNA. This fraction is not homogeneous but consists of a number of types of sequence with different lengths and repetition frequencies. In hexaploid wheat, for example, 70%–80% of the genome is made up of families of sequences present in from 100 to 100,000 copies (Smith & Flavell, 1975). The question of the composition, organization and function of these sequences is still under investigation but at least three different classes can be distinguished. These are (a) clustered sequences of repetitive DNA which are transcribed into RNA, (b) clustered sequences which are apparently not transcribed, and (c) reverse-repeats distributed at intervals throughout a substantial part of the genome.
9.6.1.1—
Transcription of Repeated Sequences
DNA-RNA hybridization has shown that a small part of the repeated sequences represents genes for rRNA. These consist of regularly arranged alternating regions of transcribed and non-transcribed DNA. The transcribed sequences can be subdivided into those where the products are stable, the 18s and 25s rRNA; and those that have metabolically unstable products, the transcribed spacer DNA in the rRNA precursor. The role of the transcribed and nontranscribed spacer sequences has not yet been unequivocally established although obvious possibilities are that they may be involved in genetic organization or regulation of transcription. Different groups of rRNA genes may sometimes be present on a number of chromosomes but as far as is known they are all practically identical.
Other families of repeated genes which have been shown to be transcribed include those for 5s RNA, tRNA and, at least in animals, the genes for histone proteins. What proportion of the genome do these account for? The actual value is higher than at first appears because hybridization only detects one strand of the DNA and normally takes no account of either the transcribed or non-transcribed spacer. This means, for example, that a 0.5% hybridization level for rRNA probably accounts for about 2.5% of the total DNA. Similar calculations for the 5s, tRNA and histone genes show that the total number of known sequences of this type only account for something like 5% of the genome.
9.6.1.2—
Palindrome Sequences
A small fraction of rapidly-renaturing DNA is known to be interspersed throughout a large part of the genome. The evidence for this comes from the 'zero-time binding' of DNA to hydroxyapatite. When DNA is denatured and immediately passed through hydroxyapatite a fraction of the DNA binds, suggesting that it is double-stranded. This happens even at very low Cot values where reassociation probably does not occur. The most convincing explanation of this behaviour is that it is due not to strand reassociation but to intra-strand 'snap-back'. This would occur if double-stranded DNA contained reverse-repeat sequences of the following type:
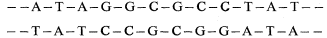
With this arrangement, termed a 'palindrome' sequence, the potential complementarity between adjoining regions of the same single-stranded DNA molecule results in rapid hairpin formation. In consequence any single-stranded DNA molecule will bind to hydroxyapatite providing it contains a double-stranded hairpin region. This provides a means of deducing the organization of palindrome sequences in the genome. The amount of DNA that binds to hydroxyapatite at zero time is markedly dependent upon the fragment size of the DNA used. This suggests that the palindrome sequences are interspersed with other regions of DNA. It has been calculated that in wheat the average distance between these palindrome sequences is 8,500 nucleotide pairs. Between 4% and 10% of the DNA is palindromic and such sequences are spread throughout at least 40% of the genome (Smith & Flavell, 1975).
There are several possible functions for such reverse-repeats. Any RNA molecule transcribed from DNA interspersed with sequences of this type would be expected to have double-stranded hairpin regions. These are known to occur in RNA but the proportion of palindromes in DNA is probably too high to be accounted for solely on this basis. One obvious property of palindromes that may have functional significance is that of symmetry. This may be important
for the recognition of particular regions of double-stranded DNA by enzymes or regulatory elements.
9.6.1.3—
Other Types of Repetitive DNA
Satellite DNAs also consist of repeated sequences. It is clear from Table 9.3 that satellites vary substantially in buoyant density, and therefore base composition, in different plants. This variation is also reflected in sequence content and repetition frequency. Although apparently homogeneous in caesium chloride the melon satellite has been shown to consist of at least two components, judged by melting experiments, caesium sulphate centrifugation and renaturation behaviour. Preliminary estimates are that the repeat lengths of the two sequences are respectively several hundred and a million or more nucleotide pairs (Bendich & Anderson, 1974; Sinclair et al., 1975). It is of interest to ask to what extent the copies within one family of repeated sequences diverge in sequence because this knowledge may provide clues about the origin and function of repetitive DNA. Indirect information on this point comes from melting experiments carried out with native and renatured satellite DNA. When similar, but not identical, sequences renature the possibility exists for the formation of duplexes with a certain proportion of mis-matched bases. For every 1% mismatching in renatured DNA the Tm is decreased by 1.0 ± 0 3°C below that of native DNA (Britten et al., 1974). Judged by this criterion the sequences within the two types of melon satellite DNA are remarkably similar. Many animal satellite DNAs appear to have evolved by duplication of a rather short sequence of DNA followed by divergence from the basic sequence. They are believed not to be transcribed. In contrast, the melon satellite DNAs appear to show very little sequence divergence and furthermore one component at least has a suprisingly large potential coding capacity. However, this may prove to be exceptional and further experimental work is necessary before it can be established whether or not plant satellites are more complex than those found in animals and whether they are ever transcribed into RNA.
The division of repetitive DNA into satellite and non-satellite components is probably an arbitrary distinction. All plants contain repeated sequence DNA irrespective of whether or not they possess satellites. This emphasises the considerable variation in properties of repeated DNA sequences. However, very little is known about their function. The fact that they sometimes form satellites suggests that they are clustered to some extent, otherwise they would remain attached to, and therefore band in association with, main-band DNA in caesium chloride. In the mouse, hybridization to cytological preparations has shown that satellite sequences are concentrated around the centromeres of many of the chromosomes, but this is not always true of satellite DNA.
Where satellites are present the amounts may vary in different cell types of the same plant (Ingle & Timmis, 1975). Furthermore there is some evidence that at least part of the repeated DNA can be dispensed with altogether. For
example, Beridze (1972) has investigated six species of Phaseolus and shown that three contain substantial amounts of satellite DNA whereas three others possess little if any satellite. Clearly, in these cases, the satellite is not essential and its presence or absence appears to have little effect on the plant phenotype. It is quite possible that at least some of the repetitive sequences in DNA are manifestations of evolutionary 'doodling'.
9.6.2—
Unique DNA Sequences
One might conclude that the non-repetitive portion of the genome represents that fraction of the DNA that codes for proteins. However, taking into account the repetitive DNA fraction, the amount of unique sequence DNA in different species is found to vary at least ten-fold (Table 9.4). One way to estimate the proportion of DNA transcribed is to carry out hybridization experiments between mRNA and unique-sequence DNA. Due to the very high Cot values required the rate and extent of hybridization using the conventional filtered technique are unsatisfactory. An alterative approach is to carry out DNA renaturation at high concentrations, using only a trace of highly radioactive RNA for hybridization (Bishop et al., 1974). However, another technical difficulty is encountered in obtaining RNA preparations sufficiently enriched with mRNA and of sufficiently high specific radioactivity for this purpose and no estimates are yet available concerning the extent of transcription of the unique fraction of the plant genome.
9.7—
RNA Synthesis
Most types of RNA are single-stranded molecules transcribed by RNA polymerase enzymes from one of the two strands of a DNA template. RNA molecules are, therefore, the direct products of genes. The following section deals with the transcription of nuclear DNA and the processing and transport of RNA to the cytoplasm. The transcription of chloroplast and mitochondrial DNA and the special case of the synthesis of RNA on a RNA template by plant viruses are excluded.
9.7.1—
General Properties of RNA
Different types of RNA are characterized by their subcellular distribution and properties such as molecular weight and nucleotide composition. Approximately 70% of cellular RNA is contained in the ribosomes where it plays a structural role. Each 80s cytoplasmic ribosome contains three different types of ribosomal RNA (rRNA). These molecules are most commonly referred to by 's values' which are related to their rates of sedimentation during ultracentrifugation.
The larger ribosomal subunit contains two rRNA molecules of 25s and 5s and the smaller subunit contains a single rRNA of 18s. The approximate molecular weights are 1.3 × 106 daltons for 25s rRNA (about 3,900 nucleotides), 0.7 × 106 daltons for 18s rRNA (about 2,100 nucleotides) and 4 × 104 daltons for 5s RNA (about 120 nucleotides). The nucleotide compositions of a range of rRNAs are shown in Table 9.6. Transfer RNA (tRNA or 4s RNA) accounts for a
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
further 25% of the total. In contrast to rRNA this fraction is heterogeneous. It consists of a range of similar but not identical molecules approximately 80 nucleotides long, each capable of being charged with a specific amino acid. By interacting with messenger-RNA appropriately charged tRNAs are responsible for aligning amino acids in the correct order during protein synthesis. In addition to the four major bases tRNA contains a very high proportion of modified or unusual bases. The most interesting type of RNA, mRNA, is present in small amounts and constitutes less than 5% of the total. It is this fraction that directs the synthesis of specific proteins by the ribosomes. The length and nucleotide
sequence of mRNAs are related to the properties of the proteins for which they code. As most plant cells synthesize a variety of different proteins the mRNA fraction generally consists of a range of molecules of different size and nucleotide composition. The fractionation, by polyacrylamide gel electrophoresis, of total RNA from suspension cultures of sycamore cells is shown in Fig. 9.13a. The rRNA and tRNA are clearly distinguished but the small amount of mRNA present is not detected. In Fig. 9.13b purified radioactive mRNA from polyribosomes of sycamore cells is compared with non-radioactive rRNA. The distance moved by each RNA through the gel is, with a few exceptions, inversely related to the log of the molecular weight (Loening, 1968, 1969).
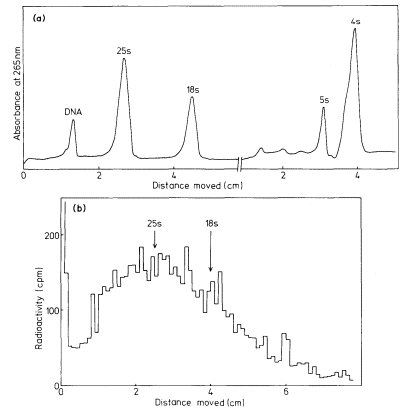
Figure 9.13
The fractionation of sycamore (Acer pseudoplatanus ) RNA by gel
electrophoresis. (a) Composite of two gel profiles. On the left the
fractionation of DNA and 25s and 18s rRNA in a 2.4% acrylamide
gel. On the right the separation of 5s from tRNA in a 7.5% gel. (b)
Gel electrophoresis of poly (A)-containing mRNA purified from
polyribosomes of sycamore cells labelled with [3 H]-uridine for
30 minutes. The positions of unlabelled 25s and 18S rRNA used
as molecular weight markers in the same gel are indicated.
(Unpublished results of D. Grierson.)
9.7.2—
Ribosomal RNA Synthesis
9.7.2.1—
The Ribosomal RNA Precursor
The DNA that codes for rRNA is known to be associated with nucleoli and the transcription of nucleolar DNA has been studied with the electron microscope (Fig. 9. 11). More detailed information has been obtained by the labelling of intact seedlings, plant segments, or cultured cells, with [32 P]-phosphate or [3 H]-uridine for various times followed by the extraction and characterization of the rapidly-labelled RNA. The mechanism of rRNA synthesis is fairly well understood and a generalized scheme outlining the main features of the process is given in Fig. 9.14. The first stable gene product which can be detected is an rRNA precursor molecule. The molecular weight of this RNA varies in different plants but is generally within the range 2.2–2.6 × 106 daltons. The rRNA precursor from mung bean roots, detected by polyacrylamide gel electrophoresis,
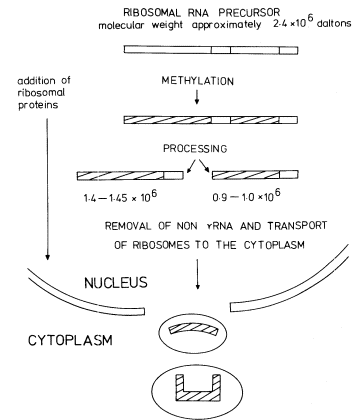
Figure 9.14
The synthesis and processing of the precursor to rRNA.
(Modified from Grierson et al., 1975.)
is shown in Fig. 9.15. The precursor undergoes considerable post-transcriptional modification in the nucleolus before it is converted into 'mature' rRNA and enters the cytoplasm. Shortly after synthesis the molecule becomes methylated at selected sites. This can be demonstrated by incubating plants in methyl-labelled methionine for a short time. The methyl groups become incorporated into the rRNA precursor via s-adenosyl methionine (Cox & Thurnock, 1973). Ribosomal-RNA is known to contain 2'-methyl ribose at certain positions. Only a small number of the residues are modified in this way and it is probable that methylation is completed during the rRNA precursor stage.
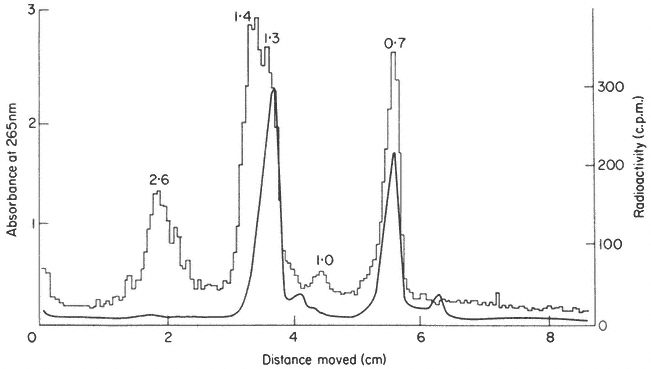
Figure 9.15
Gel electrophoresis of total RNA from roots of P. aureus labelled for
1.5 h with [32 P]-phosphate. The molecular weights of the rRNA precursor and
processing intermediates are shown in millions. (Unpublished result of D. Grierson.)
The precursor molecules correspond to the lateral fibrils attached to the matrix units of active nucleolar DNA (Fig. 9.11). Each molecule contains one sequence of 25s rRNA, one sequence of 18s (total molecular weight = 2.0 × 106 daltons) together with some non-rRNA. The evidence for this is as follows. Firstly, although it is rapidly labelled, the precursor does not accumulate in large amounts. This suggests, therefore, that it must either be rapidly degraded or converted to some other molecular form (Rogers et al., 1970; Leaver & Key, 1970; Grierson et al., 1970; Cox & Turnock, 1973; Grierson & Loening, 1974). This latter explanation would account for the fact that radioactivity appears in mature rRNA slightly later than in the precursor. Secondly, the nucleotide composition of the rRNA-precursor is similar to that of rRNA (Rogers et al., 1970; Leaver & Key, 1970; Grierson & Loening, 1974). Thirdly, competition hybridization experiments have shown that both unlabelled 25s and 18s rRNA compete with radioactive precursor-RNA for sites in the DNA, suggesting that
they share common sequences (Fig. 9.16). Finally, partial digestion of radioactive 25S, 18s and precursor RNA by T1 ribonuclease, followed by two-dimensional fractionation of the fragments shows that the 'fingerprint' pattern of the precursor is similar to that expected of a mixture of 25s and 18s rRNA (Sen et al., 1975).
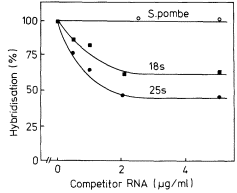
Figure 9.16
Competition hybridization between unlabelled
rRNA and radioactive rRNA precursor for
similar sites in P. aureus DNA.
(From Grierson & Loening, 1974.)
In some plants more than one precursor is detected (Leaver & Key, 1970; Cox & Turnock, 1973). The two precursors from cultured sycamore cells are shown in Fig. 9.17. In some instances the second precursor may originate in chloroplasts or mitochondria (Grierson & Loening, 1974; Kuriyama & Luck, 1973), but in carrot and sycamore both precursors are thought to be in the nucleus (Leaver & Key, 1970; Cox & Turnock, 1974). It is not clear in these cases whether the two RNAs contain similar sequences or whether they are transcribed from different genes. It is possible that the larger RNA is converted to the smaller one, but careful analysis of the rate of labelling of both molecules does not support this suggestion (Cox & Turnock, 1973; Leaver & Key, 1970).
9.7.2.2—
Processing of the Precursor
In addition to becoming methylated, and while still in the nucleolus, the precursor becomes associated with protein molecules and is processed by nucleolytic enzymes which remove the excess non-rRNA in stages to produce mature rRNA in ribosomal subunits ready for transport to the cytoplasm. Some RNA processing intermediates are present in the sample shown in Fig. 9.15. The scheme for processing outlined in Fig. 9.14 is consistent with some, but not all, the studies on rRNA synthesis in plants. For example, no immediate precursor to 25s rRNA is detectable in rye embryos (Sen et al., 1975). Furthermore the 1.0 × 106 daltons RNA is not always detected wheras certain additional RNAs
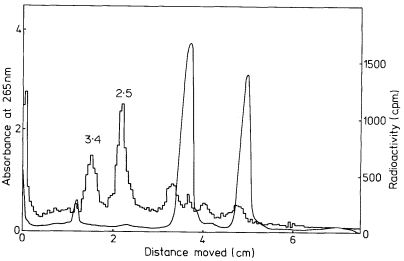
Figure 9.17
Pulse-labelled RNA from sycamore cells showing two rRNA precursor
molecules. The cells were labelled for 40 minutes with [3 H]-uridine and
total nucleic acid fractionated by gel electrophoresis. The DNA was not
removed but an identical distribution of radioactivity is observed when
this is carried out. The approximate molecular weights of the precursor
RNAs are indicated in millions. (Unpublished result of D. Grierson.)
are sometimes observed. For example, after cycloheximide treatment of cultured parsley cells, which slows down rRNA processing, previously undetected RNAs with molecular weights of 2.0 and 0.9 × 106 daltons were found (Gebauer et al., 1975). This suggests that if certain processing events occur very rapidly they may normally go undetected. It is very probable that the complete sequence of processing steps will only be worked out when it becomes possible to study accurately RNA metabolism in isolated nucleoli. Apparent differences between closely related species may be explained by assuming that certain critical stages of processing occur at different rates. In addition, it should be realised that the processing enzymes are functioning at different, possibly distant, sites within the same molecule. There seems no compelling reason to expect these modifications always to occur in the same chronological order and, if they do not, variations in the pattern of processing intermediates might be expected under altered conditions or in different species.
The function of the excess RNA in the precursor is not known. It is too variable in size in different organisms to be a mRNA for the ribosomal proteins (Loening, 1970) and in any case it appears to be degraded very rapidly after synthesis. It may play some role in the regulation of transcription but more probably it is necessary for ensuring that rRNA adopts the correct secondary and tertiary structure required for methylation and for the addition of proteins during ribosome assembly.
It is pertinent to ask whether the multi-stage mechanism that operates for rRNA production is unique or whether it represents a general principle common to the synthesis of all types of RNA. Precursors to tRNA and 5s RNA have been identified and characterized in bacteria and animals (Hecht et al., 1968; Bernhardt & Darnell, 1969) and precursors of mRNA have also been detected, although there is uncertainty about the actual details of processing (Brawerman, 1974). Although RNA metabolism has not been so intensively studied in plants the indications are that precursors to 5s RNA, tRNA and mRNA do occur. When considering the question of the regulation of RNA metabolism, therefore, it is necessary to take account of post-transcriptional modifications as possible control points.
9.7.3—
Messenger-RNA Synthesis and Metabolism
The question of mRNA synthesis has proved extremely difficult to investigate due largely to problems of identification and purification of mRNA. In the past, rather imprecise criteria for mRNA identification, such as size heterogeneity, similarity in nucleotide composition to DNA, rapidity of labelling and association with polyribosomes, were adopted. More recently, two additional approaches have become available. Firstly, the development of an in vitro protein synthesizing system has made it possible to test the capacity of an RNA fraction to direct the synthesis of specific proteins. Secondly, the discovery that many mRNAs contain sequences of poly (adenylic) acid (poly (A)) has provided a simple approach to the purification of mRNA.
9.7.3.1—
Poly (A)-Containing RNA
It has been known for some considerable time that a small fraction of total plant RNA has a very high AMP content. This fraction, often referred to as DNA-like RNA or D-RNA, is considered to consist, at least in part, of mRNA and is partially purified by chromatography on columns of methylated albumin adsorbed to kieselguhr (Ingle et al., 1965; Lin et al., 1966; Jackson & Ingle, 1973). More recent work with animal mRNA has shown that the high AMP content is due to the presence of regions of poly (A) in the mRNA molecules. These sequences of poly (A) are generally 50 to 250 nucleotides in length. In animals they are attached to the 3' end of the mRNA molecules after transcription (Brawerman, 1974) although there is some evidence that shorter regions of poly (A) may also be introduced into RNA as part of the transcription process (Jacobson et al., 1974). Poly (A)-containing RNA has been identified in a variety of animals (Brawerman, 1974; Matthews, 1973) and plants (Sagher et al., 1974; Verma et al., 1974; Higgins et al., 1973) although it has not been found in higher plant chloroplasts (Wheeler & Hartley, 1975). It is apparently present in mitochondria from some organisms (Perlman et al., 1973; Avadhani et al.,
1973; Ojala & Attardi, 1974) but not others (Groot et al., 1974). This may reflect true differences between organisms or alternatively may be a question of the experimental technique. For example, contrary to earlier reports that poly (A) is absent from mRNA from bacteria, it has recently been shown to be present at a low level in E. coli. (Nakazato et al., 1975).
The function of the poly (A) sequences is not known but their presence can be used to purify mRNA. There are a number of methods for doing this. Poly (A) will bind to nitrocellulose or cellulose at high salt concentrations. RNA without poly (A) can be washed off and the poly (A)-containing RNA is then eluted at low ionic strength. However, cellulose and nitrocellulose do not necessarily retain all the poly (A)-containing RNA. A more specific type of binding is achieved using columns of oligo (deoxythymidylic) acid attached to cellulose or poly (uridylic) acid attached to sepharose. Both these polymers rapidly form stable nucleic acid hybrids with the poly (A) sequence attached to the mRNA. Other types of RNA pass straight through the column, although it is sometimes necessary to wash through extensively with a buffer. The poly (A)-containing RNA is only released after lowering the salt concentration or raising the temperature to cause melting of the hybrid between poly (A) and either oligo (dT) or poly (U).
There is considerable evidence that poly (A)-containing RNA from plants is mRNA. It is associated with polyribosomes (Higgins et al., 1973; Grierson et al., 1976) and its synthesis is inhibited by the antibiotic a -amanitin which prevents transcription of non-rRNA in the nucleus (Higgins et al., 1973). When purified from cells synthesizing a large number of proteins of different sizes poly (A)-containing RNA is polydisperse, but when extracted from soybean root nodules, which synthesize substantial amounts of leghaemoglobin, two peaks of RNA are detected, each presumably consisting of similar or identical RNA molecules. These RNA fractions stimulate protein synthesis by a cell-free system from wheat germ. Furthermore, leghaemoglobin has been positively identified among the products of in vitro synthesis (Verma et al., 1974). Supporting evidence that poly (A)-containing RNA stimulates protein synthesis comes from a large number of studies and the small number of individual proteins identified as in vitro products is growing steadily. However, it should be stressed that not all mRNA molecules contain poly (A). In animals, for example, about 70% of mRNA is estimated to contain poly (A) (Milcarek et al., 1974) whereas only about 40% of polydisperse RNA from polyribosomes of cultured sycamore cells can be purified by oligo (dT)-cellulose (Covey & Grierson, 1976). The functional significance of the two mRNA populations is not clear.
9.7.3.2—
Postranscriptional Modification of mRNA
Poly (A)-containing RNA is synthesized in the nuclei of higher plant cells. Total rapidly-labelled RNA from nuclei consists of the rRNA precursors and pro-
cessing intermediates together with polydisperse RNA (Fig. 9.18). Approximately 15% of this RNA is retained by oligo (dT)-cellulose and by this criterion is judged to contain poly (A) sequences (Grierson & Covey, 1976). It is polydisperse in size and has a high AMP content (Table 9.6). There is little direct evidence concerning the mechanism of poly (A) addition to nuclear polydisperse RNA in plants. It is assumed to be added to the 3' end of the molecules by the enzyme ATP-poly-nucleotidylexotransferase after transcription by RNA polymerase (Maale et al., 1975). The poly (A) sequences in higher plant RNA have been estimated to range from 50–250 nucleotides in length (Sagher et al., 1974; Covey & Grierson, 1976). However, plant DNA appears to contain short sequences of AT base pairs which could give rise to short poly (A) regions by transcription (Grierson, 1975) and there is evidence that similar regions are transcribed in slime moulds (Jacobsen et al., 1974).
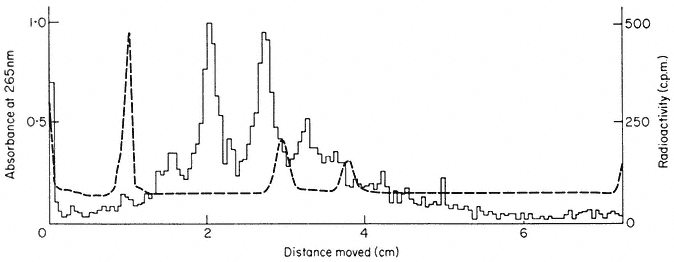
Figure 9.18
Gel electrophoresis of total nucleic acid from nuclei of sycamore cells.
The cells were labelled for 1 h with [3 H]-uridine, a nuclear fraction was
prepared and total nucleic acid extracted and fractionated by gel
electrophoresis. The radioactivity is shown as a histogram.
Note that the nuclear fraction contains very
little rRNA by comparison with Fig. 9.17.
(Grierson & Covey, 1976.)
Comparison of the nuclear and cytoplasmic poly (A)-containing RNA by gel electrophoresis suggests that nuclear RNA is reduced in size before entering the cytoplasm (Fig. 9.19). This suggests that there may be a processing mechanism analogous to that which operates for rRNA synthesis. In slime moulds the nuclear precursor appears to be approximately 20% larger than cytoplasmic mRNA. (Firtel & Lodish, 1973). In higher plants the average molecular weight of the nuclear poly (A)-containing RNA is 1 × 106 daltons (Grierson & Covey, 1976). This is in contrast to the giant precursors to mRNA earlier claimed to occur in animal cells. Very little is known about the processing mechanism, but Firtel and Lodish (1973) have shown that the nuclear precursor contains a sequence of about 300 nucleotides near the 5' end of the molecule which is lost before the RNA enters the cytoplasm. This is of considerable interest
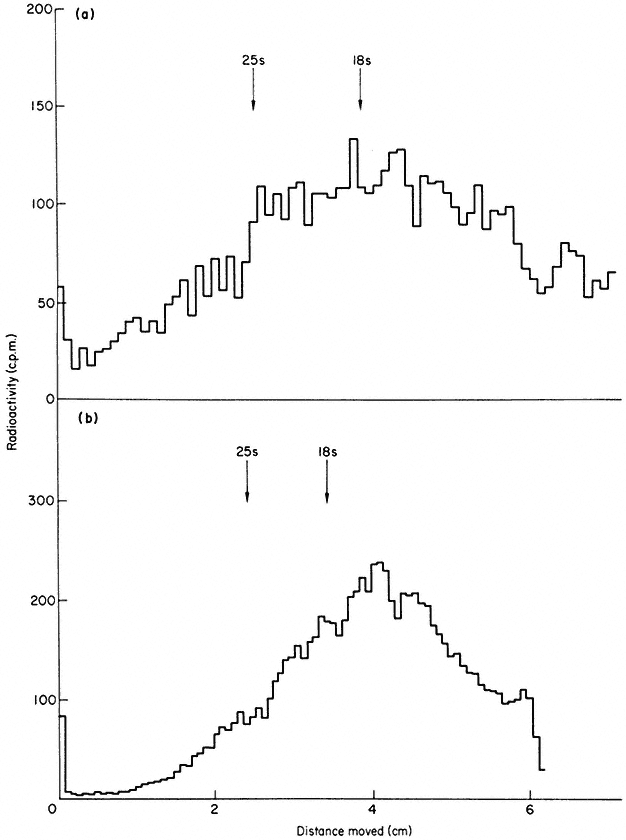
Figure 9.19
Gel electrophoresis of poly (A)-containing RNA from nucleus and cytoplasm.
Pulse labelled sycamore cells were separated into (a) nuclear, and (b) cytoplasmic
fractions and poly (A)-containing RNA purified by oligo (dT)-cellulose chromatograph.
The two RNA preparations were denatured in 8 B urea at 60°C to disrupt aggregates
and fractionated by gel electrophoresis in the presence of marker rRNA. The cytoplasmic
RNA is distinctly smaller, on average, than the nuclear RNA. (Grierson & Covey, 1976.)
because this region appears to be transcribed from repetitive DNA. It therefore seems that some of the repeated DNA sequences interspersed between nonrepetitive DNA are transcribed and may play a regulatory role in processing or transport of mRNA.
9.8—
Nucleic Acid-Protein Interactions
9.8.1—
Histones
Chromosomal DNA is invariably associated with several different types of proteins (Table 9.7). These include histones which occur in most eukaryotes.
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
There are five main histone proteins designated F1 , F2b , F2a1 , F2a2 and F3 , with molecular weights from 11,000 to 22,000 daltons. They are strongly basic and generally have a high content of arginine and lysine. The basic amino acids are often clustered towards either the N or C terminus of the proteins (Macgillivray & Rickwcod, 1974) suggesting that these regions interact directly with the DNA. Stedman and Stedman (1950) proposed that histones are responsible in some way for the regulation of gene expression but it now seems that they lack the specificity for recognition of different gene sequences and it is more likely that they fulfil a more general role in the organization of chromosomes. Comparison of the amino acid sequences of histones from different species suggests that they have been highly conserved during evolution. For example, the sequences of F2a1 from calf, pig and rat are identical and differ only in two substitutions from F2a1 from pea (Delange et al., 1966). This suggests these proteins evolved to fulfil an essential role with strict requirements
for a precise sequence, although other histones, notably F2b and F2a2 show greater variability. Histones frequently undergo modifications such as methylation, acetylation or phosphorylation. These are normally reversible and would be expected to alter the association of histones with DNA. In general these modifications occur at about the time of DNA synthesis although they may also herald an increase in RNA synthesis. During the cell cycle the synthesis of histones is closely linked to that of DNA and phosphorylation of F1 has been shown to precede cell division (Bradbury et al., 1974). Basic non-histone proteins and acidic proteins also occur in association with DNA. These form a more heterogeneous group than the histones and include components of different molecular weight, charge and tissue distribution. They are less stable metabolically than histones and do not behave as a single class of proteins. Several lines of evidence suggest that this group may include regulatory elements (Macgillivray & Rickwood, 1974).
9.8.2—
Subunit Structure of Chromatin and Chromosome Organization
The double helical structure of DNA is widely known and it is equally widely recognized that a further degree of organization is responsible for the folding of 1 m or more of DNA into each set of chromosomes. Early observations on the structure of chromosomes indicated that DNA is organized in fibres of the order of 10 nm thick and it was suggested that the double helix, in association with proteins, is supercoiled. In addition, X-ray studies suggested that there is a repeating structure in chromatin at intervals of approximately 10 nm and more recently a linear arrangement of beads of the order of 7 nm diameter has been observed in chromatin with the aid of the electron microscope (Olins & Olins, 1974). Further evidence for a regular arrangement of DNA in chromatin comes from the observation that following nuclease digestion reproducible fragments of DNA approximately 205 base pairs long are produced (Noll, 1975). Kornberg (1974) has suggested a repeating subunit structure of approximately 200 DNA base pairs associated with 2 molecules of each of the four major histones (F2b , F2a1 , F2a2 and F3 ) and one molecule of F1 . In this model the DNA double helix is envisaged to be folded to approximately one seventh its length and Crick and Klug (1975) have suggested that this could occur by 'kinking' of the DNA helix at intervals of perhaps 20 base pairs. Bradbury and his colleagues (Baldwin et al., 1975) have proposed that repeating units of DNA approximately 200 base pairs long are coiled on the outside of aggregates of the four major histones which associate with each other by apolar interactions, leaving their basic regions free to associate with the DNA. Histone F1 , which is believed not to be directly involved in the subunit, may perform a cross-linking function. Only about 87% of the DNA in chromatin is protected against deoxyribonuclease digestion (Noll, 1975) and it is possible that not all the DNA participates in formation of this type of subunit.
Further insight into the functional organization in chromatin comes from studies on the giant chromosomes from certain insects. These chromosomes consist of a series of densely coiled bands connected by less dense regions. These are much larger than the repeating unit of 200 DNA base pairs. Each band plus interband region is considered to correspond to a gene because there is good numerical agreement between their number and the number of complementation groups determined by breeding experiments. However, calculations show that only a fraction of the DNA in each band plus interband region is necessary in order to code for a protein (Beerman, 1972) and the question of the function and organization of the DNA remains unresolved. Regions of the DNA being actively transcribed appear as chromosome puffs composed of DNA, RNA and protein. Crick (1971) has suggested that the interband regions contain DNA that is transcribed and that each band consists of regulatory DNA sequences. In contrast, Paul (1972) has proposed a different type of arrangement.
Different regions of chromosomes often contain chromatin condensed to different extents (Macgillivray & Rickwood, 1974). Euchromatin appears to be loosely packed. It is known to contain both unique and repeated sequences and is actively transcribed. Heterochromatin is more tightly packed and consists of two types. Constitutive heterochromatin contains sequences that are never transcribed and includes repeated sequences. In some species satellite DNAs are known to be clustered in heterochromatic regions near the centromeres. Finally, facultative heterochromatin constitutes those regions of the chromosome that sometimes appear as euchromatin.
9.8.3—
Regulation of Transcription
Thus there is evidence for the coiling of DNA and formation of regular DNA-protein complexes which may be further folded into larger units. Different regions of the chromosome are folded to different extents and this presumably affects the availability of DNA for transcription. Comparison of the extent of transcription of pure DNA and isolated chromatin by added bacterial RNA polymerase suggests that in animals only about 10% of the genome is available for RNA synthesis. Hybridization experiments suggest that the RNA synthesized from isolated animal chromatin is tissue specific and resembles in sequence the type of RNA synthesized in vivo (Macgillivray & Rickwood, 1974). It is probable that the non-histone proteins are more important in determining which segments of the genome are repressed but further characterization of the properties and mechanism of action of these proteins is necessary before their function becomes clear (see Mcgillivray & Rickwood, 1974). It has also been suggested that chromosomal RNA may be important in regulating gene expression, possibly in conjunction with repeated sequences in DNA (Georgiev, 1969; Ryskov et al., 1971; Britten & Davidson, 1969). There is undoubtedly RNA associated with chromatin, including growing chains of polydisperse-
RNA and rRNA-precursor, but the existence of an additional type of regulator RNA has been proposed. However, at the present time there is a certain amount of controversy concerning the characterization and properties of chromosomal RNA (Jacobson & Bonner, 1971; Huang, 1967; Heyden & Zachau, 1971; Holmes et al., 1972). As yet there is no evidence that this RNA regulates transcription.
In addition to repression of the genome by interaction with proteins, template selection may also be controlled by RNA polymerase enzymes and by other protein factors that interact with the polymerases to regulate transcription. As far as studies on plant chromatin are concerned in vitro RNA synthesis has been achieved using the endogenous RNA polymerases that remain attached to the isolated chromatin (Guilfoyle & Hanson, 1974). However, there have been few attempts to characterize the RNA products in detail. There are several distinct RNA polymerases with different properties. These enzymes are probably responsible for transcribing different regions of the genome. For example, RNA polymerase I, which is insensitive to the antibiotic a -amanitin, is located within the nucleolus (Brandle & Zetsche, 1973). This enzyme appears to be responsible for transcription of the rRNA genes. The activity of this enzyme and the rate of RNA production are significantly enhanced in tissue treated with auxin (Guilfoyle et al., 1975). Enzymes sensitive to a -amanitin are present in the nucleoplasm and presumably are responsible for transcribing other parts of the genome. These polymerases may also be affected by hormones via the mediation of hormone-binding factors (see chapter 13). In addition, evidence has been obtained for the existence of other protein factors which govern the activity of RNA polymerases by stimulating initiation of RNA synthesis (Mondal et al., 1972a, 1972b).
9.9—
Conclusions
Now that we have a general idea of the organization and properties of DNA it is possible to pose more precise questions about the function of the different parts of the genome. For example it would be interesting to establish what proportion of the DNA is transcribed into RNA in plants with different nuclear DNA contents. There is also the question of the function of the repeated DNA sequences: are they involved in regulation of transcription; to what extent are they transcribed; and what is the function of the repetitive sequences in the nuclear precursors to mRNA? Further characterization of the various RNA polymerase enzymes and the protein factors which influence RNA synthesis will doubtless increase our knowledge of the control of gene expression. However, it does seem that ultimately such studies should be carried out using chromatin as template in order to understand the precise molecular interactions involved. Finally, the possibility of post-transcriptional regulation should not be overlooked. It is important to establish the role of post-transcriptional modification
of RNA molecules in relation to mRNA selection and transport to the cytoplasm.
Further Reading
Biswas B.B., Ganguly A. & Das A. (1975) Eukaryotic RNA polymerases and the factors that control them. In Progress in Nucleic Acid Research and Molecular Biology (ed. W.E. Cohen), Vol. 15, pp. 145–84. Academic Press, New York.
Bryant J.A. (1976) Nuclear DNA. In Molecular Aspects of Gene Expression in Plants (ed. J.A. Bryant), pp. 1–51. Academic Press, London.
Duda C.T. (1976) Plant RNA polymerases. Ann. Rev. Pl. Physiol.27, 119–32.
Grierson D. (1976) RNA structure and metabolism. In Molecular Aspects of Gene Expression in Plants (ed. J.A. Bryant). pp. 53–108. Academic Press, London.
Kornberg A. (1974) DNA Synthesis. W.H. Freeman & Co., San Francisco.
Lewin B. (1974) Gene Expression, Volume 2, Eukaryotic Chromosomes. J. Wiley & Sons Ltd., London.
Nagl W. (1976) Nuclear organisation. Ann. Rev. Pl. Physiol.27, 39–69.
Parish J.H. (1972) Principles and Practice of Experiments with Nucleic Acids. Longman, London.
Rae P.M.M. (1972) The distribution of repetitive DNA sequences in chromosomes. In Advances in Cell Biology and Molecular Biology (ed. E.J. Dupraw). Vol. 2, pp. 109–49. Academic Press, New York.
Stewart P.R. & Letham D.S. (eds.) (1973) The Ribonucleic Acids. Springer-Verlag, Berlin.